23 May 2026
Manish Kumar

Everyone talks about LLMs, but the real magic is often in the pieces around them: tokens, embeddings, retrieval, adapters, and compressed weights. This post gently opens the black box and shows how these ideas fit together to power the AI tools we use every day - without drowning you in jargon.
More …
11 Apr 2023
Manish Kumar

Unless you’re living under a rock, you probably have heard about ChatGPT by now. The Internet has been buzzing with new technologies for the past few months. Tools that would have taken months, possibly years, to build are being developed just over a weekend of hacking. I truly believe that we’re living in one of the most exciting times in history.
More …
22 Apr 2021
Manish Kumar

Let’s try to get a brief overview of what UMLS is, and then we’ll jump into more details. So, in simple terms, UMLS is a collection of many controlled biomedical vocabularies to provide cross-walk among them and to enable interaction between computer systems. To simplify it further, there are multiple biomedical vocabularies out there developed by different organizations. Each of them has come up with their own set of IDs assigned to each term. UMLS helps in mapping each of them to one common ID.
More …
11 Apr 2021
Manish Kumar
This is a continuation of a 2 part series in predicting the products of a chemical reaction using deep learning methods. Checkout the first part here.
In part 1, we:
- First introduced the problem statement
- Then we saw how a molecule is represented in a graph network
More …
10 Apr 2021
Manish Kumar
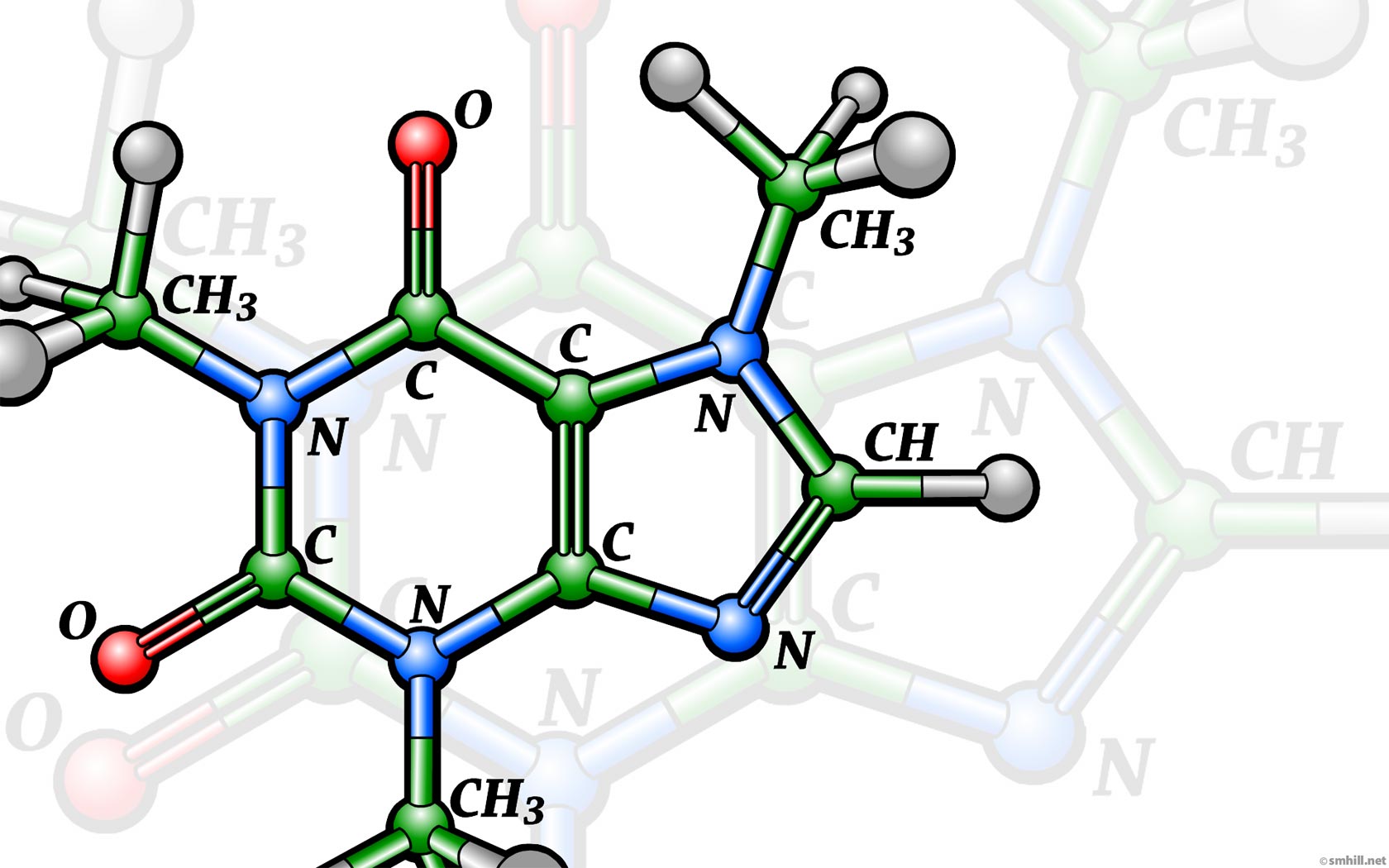
Reactant A and B reacts to form the products C and D. The chances are that you have encountered this type of chemical reaction in your school days. While on paper they may seem as simple as drawing a bunch of arrows, in the real world, even the simplest of those reactions, takes a long time, money, and resources to formulate. The reaction time can vary from the order of seconds to the order of months.
More …

 ChemicBook
ChemicBook



